|
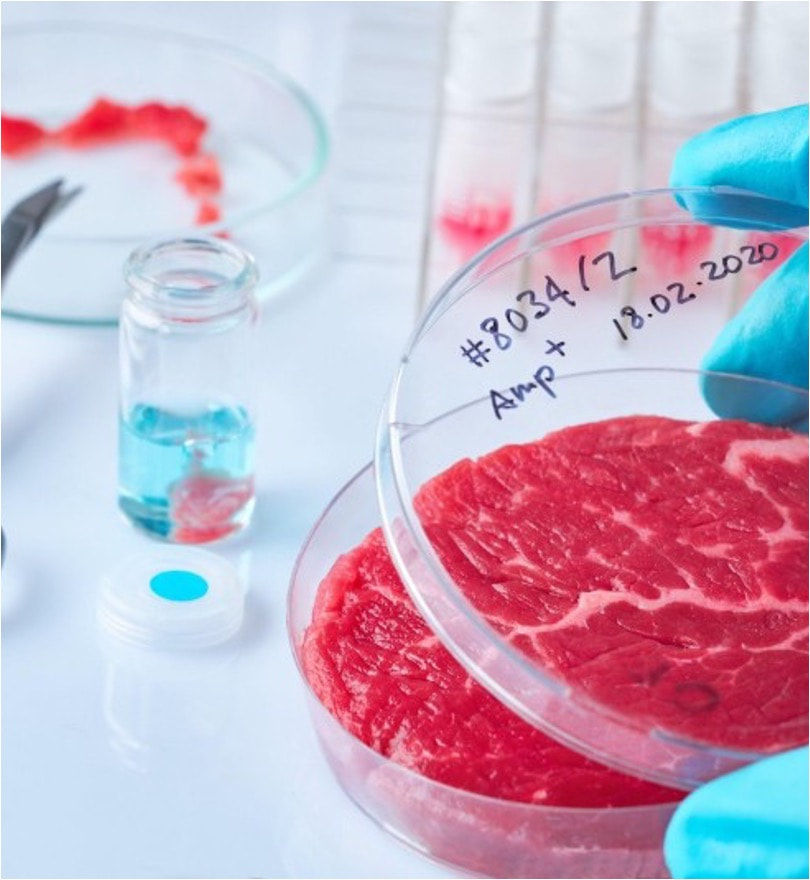
di Alessandro Bertero Negli ultimi mesi in Italia si è sentito molto parlare di "carne sintetica" - che di sintetico non ha nulla e sarebbe appropriato chiamare "carne coltivata". Nel mio piccolo ho cercato di fare chiarezza con articoli, interventi radiofonici (i.e. radio capital - 02:45:45; radio 1 - 41:30), interviste (i.e. Le Scienze; Servizio Pubblico), e seminari online (i.e. Associazione Luca Coscioni; Partito Democratico Stati Uniti). Purtroppo, questi ed altri contributi di colleghe e colleghi attivi nel settore non hanno dissuaso il Senato della Repubblica ad approvare lo scorso 20 Luglio il disegno di legge s.651, che si pone l'obiettivo di vietare le attività commerciali nel settore della carne (vertebrata) coltivata in Italia. In questa fase non ci fu molta occasione di fare chiarezza direttamente con i Senatori chiamati a votare la norma in quanto vennero solo richiesti dei contributi scritti. La Sen. Cattaneo tentò di intavolare tale discussione con un ottimo convegno in merito, ma nè questo nè gli eccellenti interventi della Sen. Cattaneo e di altri esponenti dell'opposizione (i.e. Sen. Floridia, Unterberge, Zambito, e Maiorino) cambiarono un esito scontato dal punto di vista politico. L'unico passo in avanti fu la rimozione della nomenclatura "cibo sintetico" dal ddl, una magra consolazione. Il disegno di legge è ora in discussione alla Camera dei Deputati, nello specifico all'esame delle Commissioni riunite Affari Sociali ed Agricoltura. Questa volta ho apprezzato l'invito a contribuire ad una audizione informale insieme ad altri colleghi esperti in materia a vario titolo. E' stata un'ottima occasione di confronto, anche tra gli auditi, e che speriamo possa aiutare a far chiarezza sul tema e, siccome la speranza è l'ultima a morire, un ripensamento sul ddl. Per chi fosse interessato ecco il link alla registrazione (inizio delle audizioni ad esperti in materia al minuto 22): https://webtv.camera.it/evento/23243# Per coloro che arrivassero al termine dell'audizione alcuni chiarimenti sul dibattito tra uditi che non è stato possibile completare in diretta.
In primis, nel mio secondo intervento menzionai la statistica secondo la quale 9 calorie di cibo servano per ottenere 1 caloria di carne di pollo per essere poi corretto dal Prof. Cocconcelli, che riportò come la food conversion rate sia di 1.2. Siccome questo aspetto esula dal mio campo di ricerca mi ero affidato al Good Food Institute per informarmi in merito, ed ho quindi approfondito per comprendere come mai le due statistiche fossero così divergenti. Capisco ora che la confusione nasca dal come si calcola la conversione di cibo: se si considera il peso in kg e l’animale intero allora il rapporto è effettivamente più vicino a quello citato dal Prof. Cocconcelli (anche se mi risulta più alto, 1.77 al 2022 https://www.nationalchickencouncil.org/about-the-industry/statistics/u-s-broiler-performance/ ); se si considera invece il potere calorico del mangime e delle carne allora il rapporto è in linea con la statistica del GFI che citai (7.7, ovvero 13% di efficienza di conversione, vedasi Tabella 1 di questo studio del 2016). Lo stesso studio chiarisce come la "food conversion ratio" citata dal Prof. Cocconcelli diventi molto più alta qualora si consideri solo il peso edibile dell’animale (4.2 ± 0.8) ed ancora di più considerando quello consumato (5.4 ± 1.4). Detto tutto ciò, è chiaro che resti da valutare l’effettivo contenuto nutrizionale della carne coltivata risultante dall’assorbimento intestinale e della biochimica cellulare, che ovviamente non consumano tutte le calorie ingerite ma solo una loro frazione. In secunids, per quanto riguarda l’impatto della produzione di amino acidi ricombinanti, ho ricontrollato i parametri della life cycle assessment (LCA) che ho citato, e come mi sembrava di ricordare viene ipotizzato l’uso di idrolizzato di soia come ingrediente fondamentale, poi integrato da alcuni amino acidi ottenuti per fermentazione/sintesi (quali la glutamina). Chiaramente queste sono per ora congetture da verificare, ma è altrettanto ovvio che se non si raggiungessero questi obiettivi tecnico-scientifici il costo del prodotto rimarrebbe non competitivo dal punto di vista economico e quindi automaticamente fuori discussione.
0 Comments
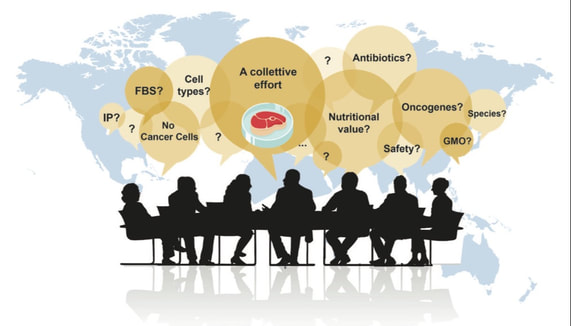
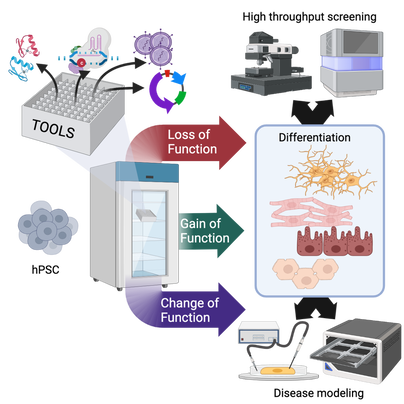 FEBS letters published our comprehensive review of "do's and don't" when manipulating and studying gene function in human pluripotent stem cells (hPSCs). We took this chance to sit down for an interview with FEBS Network to highlight our lab's favourite technique: genome editing in hiPSCs. Both of these contributions are linked to our lab's FEBS Excellence Award. We are very grateful to FEBS for these opportunities that go beyond the (extremely valuable) funds to our group! Our mammoth review (26 pages + 279 references!) involved all lab members, each covering the techniques they are more expert in (a collective experience of >30 research years!). We put particular attention to covering not only the state of the art but also the "dark knowledge": negative results and challenges that rarely find space in the published literature. We hope that this resource will help newcomers to the field to avoid hitting roadblocks as they "hitchhike" their way to altering gene function in hPSCs. Our lab has been awarded two grants from the Italian Ministry for University and Research (MUR) as part of the "Progetti di Rilevante Interesse Nazionale" (PRIN) program. As part of the PRIN 2022 PNRR call, our project entitled: "From stem cells to algae and back: circular biotechnology for cellular agriculture and biomass production" was ranked 4th overall in the LS9 scientific sector, and 1st among the "Under 40" category. This project is led by our lab and involves an exciting interdisciplinary collaboration with Alberta Pinnola's plant physiology group at the University of Pavia. The total budget is € 230,861. In this proposal we tackle a pivotal challenge towards the pursuit of a sustainable and circular-economy-based future: how can we provide food within planetary limits for an expanding population with a growing appetite for meat and fish? This is our first research grant in the cellular agricolture space, which we are super excited about. As part of the PRIN 2022 call, our project entitled: "Decoding and leveraging the molecular determinants of myogenic fate through integrative genomics and cell engineering" received a score of 83/100 in the LS2 scientific sector. The project is led by Davide Cacchiarelli's integrative genomics lab at TIGEM and involves the collaboration of our group as well as Annamaria Carissimo's bioinformatics team at CNR. The total budget is € 199,628. The overarching goal of this proposal is to first uncover and then exploit the gene regulatory networks behind cell fate conversion into muscle. The knowledge generated by this proposal will enable large scale, cost-effective generation of mature muscle cells for various applications. These two grants support our emerging lines of research in cardiac regeneration and cellular agriculture. Both grants have a two year duration and will start in the Fall of 2023. We are looking for postdocs motivated to develop these exciting lines of research, so please spread the word and reach out if interested. Nature Biotechnology published our correspondence entitled "A call for an ‘Asilomar’ for cultivated meat and seafood". As biotechnologists who are experiencing the attempts of the Italian government to ban cultivated meat and seafood, we humbly call for action from all stakeholders: either we self-regulate and build consensus on critical issues to reassure both the public and governments or history may repeat itself with prohibitions that could stifle this emerging field.
We had previously debunked the fallacious arguments brought up by detractors of cultivated meat with a comment published on Nature Italy (Italian version), so as to promote clarity on the topic at the national level. With this new perspective article we wish to bring attention to the situation at the international level, and to propose an actionable framework to prevent similar debacles elsewhere. We are ecstatic to announce that our ERC Starting Grant 2022 project "TRANS-3" (TRANscriptional and Splicing Topological Regulations Architecting Nuclear Structure in TRANS) was selected for funding. Our project was awarded a budget of ~€1.77M and will be developed over a period of 5 years. We are eager to get started and we are looking for motivated postdocs to join us - check out the job ad below and kindly help us spread the word! We have been working towards this goal for many years now, and we could not have reached it without the support of mentors, colleagues, and friends that dedicated their time and energies to help us build our most exciting project yet. For those not in the know, ERC Starting Grants are awarded to scientists with 2-7 years of research experience since their PhD. In the 2022 call only ~13,9% of nearly 3,000 applications could be supported: less than 30 of these are for projects that will be developed in Italy, and only 7 of these are in the life sciences arena. Our proposal lies in the so-called Life Sciences 1 (LS1) panel: "Molecules of Life: Biological Mechanisms, Structures and Functions", which encompasses the disciplines of molecular biology, biochemistry, structural biology, molecular biophysics, synthetic and chemical biology, drug design, innovative methods and modelling. This has been a historically underrepresented panel for scientists in Italy, which previously secured less than 2.5% of all LS1 proposals. We are thus especially proud to represent this important arena, which is essential for innovation in our country and the whole of Europe. Below are the title and abstract of our project, followed by a narrative description of our motivations and goals. Project Title Beyond the chromosome: unravelling the interplay between inter-chromosomal genome architecture and mRNA biogenesis Project Abstract In TRANS-3 we will develop the theoretical and experimental framework to understand inter-chromosomal genome structure and activity. Despite advances in sequencing chromosomes and mapping their three-dimensional (3D) organization, a full picture of 3D genome structure that details how the borders of various chromosome territories functionally interface one another is still missing. As an analogy, our world map does not indicate most natural corridors or manmade infrastructure that connect countries to one another. Worse, we barely understand how such connections function. My recent work has identified one of the few functional inter-chromosomal (trans) DNA interactions known to date: a splicing factory involving over ten chromosomes and orchestrated by the muscle-specific protein RBM20 around its key target, the TTN pre-mRNA. I hypothesize that this exemplifies how mRNA biogenesis instructs the formation of trans-interacting chromatin domains (TIDs) around mRNA factories, nuclear compartments that facilitate gene regulation. We will test this general hypothesis by dissecting the mechanisms and function of a specific, disease-relevant model: the RBM20 mRNA splicing factory. We will then explore the global impact of these regulations. First (WP1) we will mechanistically assess how mRNA factories form and act through live imaging of the nuclear positioning and alternative splicing dynamics of an RBM20-regulated locus. Secondly (WP2) we will examine the physiological role of mRNA factories by studying the effects of disease-associated mutations in RBM20 and the TTN regulator GATA4, and of genetic variability in TTN regulatory regions. Finally (WP3) we will develop and deploy a novel pipeline to identify, validate, and study new TIDs and mRNA factories through the combination of molecular biology, bioinformatics, biochemistry, and single cell biology. In all, TRANS-3 will venture beyond the chromosome frontier towards a deeper understanding of nuclear structure-function. The Heart of the Matter: How the Structure of DNA Influences Heart Disease
The human body is composed of over two hundred types of cells, each as different from one another as the heart, blood, and brain. And yet, each of the one hundred trillion human cells shares the same genetic code. How can we reconcile this apparent paradox? In a word: structure. The 46 DNA molecules that make up the human genetic code - the so-called chromosomes - are about two meters long in total. And yet, they are jealously preserved in a structure, the cellular nucleus, no larger than a hundredth of a millimeter - another paradox? In reality, this is possible because the DNA filament is extremely thin - about two millionths of a millimeter - and can therefore be folded upon itself. Anyone who has had the unpleasant experience of winding a ball of yarn knows how difficult it can be. Now imagine having to do it for 46 balls of yarn that have to intertwine with each other in a limited space. To complicate matters, DNA cannot simply be all compacted upon itself: this would make it useless! Some portions of the chromosomes, the so-called genes, which are necessary for cellular function, must be left free. To top it off, there are over 200 types of patterns to follow in folding human DNA, as many as the types of cells that make up our body. This is where the answer to our opening question lies: each cell only leaves unfolded a portion of its genetic code, namely the portion that contains the genes that characterize the function of that cell, such as the contractile proteins of muscle, the hemoglobin of blood, or the neurotransmitters of the brain. In the last twenty years, we have made tremendous strides in understanding human DNA. In addition to determining its sequence, we have mapped how the various chromosomes are folded in many different cell types. However, we are missing a fundamental detail: how are the various chromosomes packaged together? To make an analogy, it's like having detailed maps of Italy and other European countries - complete with cities and other points of interest, and the infrastructure that connects them such as roads, railways, and navigable rivers - but having no idea how to cross national borders. It would be quite difficult to get from Turin to Paris if we didn't know about the existence of the Sandro Pertini airport, the Frejus tunnels or Mont Blanc, or the various mountain passes and valleys: we would risk getting lost for weeks in the Alps and end up retreating! The purpose of our "TRANS-3" project, funded by the European Research Council (ERC), is to launch an exploratory expedition beyond the borders of individual chromosomes, to map their connections and understand their function. This information will allow for a better understanding of how the nucleus structure is altered by various diseases, and could lead to new drugs capable of curing them. Our expedition starts from the discovery of the first example of how chromosomes are not simply packed into the nucleus like socks in a messy teenager's drawer, but can develop specific and important connections. We have identified a type of DNA folding that involves ten chromosomes and occurs only in muscle cells such as those in the heart. This structure brings together a series of genes essential for cardiac function and all regulated by a protein exclusively expressed in muscle and called RBM20. Like in an assembly line, we believe that gathering the necessary machinery and components for the expression of these genes increases their efficiency. Furthermore, we know that mutations in both the RBM20 protein and its main regulatory target, the gene encoding the titin contractile protein, lead to severe cardiac diseases characterized by rhythm disorders and insufficient blood supply to the body. Therefore, we aim to examine in detail the functioning of the RBM20 "factories" both under normal conditions and following mutations that alter their function. These studies will use genetic engineering, super-resolution microscopy, genomic sequencing, and bioengineering methodologies applied to stem cells differentiated into cardiac muscle. Starting from the outpost of RBM20, we plan to explore the rest of the boundaries between the various chromosomes in heart cells. Similar to space exploration, this ambitious mission will require us to develop new technologies. In our case, we will optimize methods for mapping interactions between DNA molecules, analyzing data from a statistical point of view, identifying the proteins and genes involved, and finally validating the function of these linking structures. It will be a difficult journey, but we have five years ahead of us, a good budget, a close-knit team, and important international collaborations. Most importantly, we are armed with the motivation that even small successes in this frontier field could have important and partly unpredictable implications for understanding and treating not only cardiac diseases but also others. ------ Thank you for reading through the end; stay tuned for regular updates on our project!
This epic tale begins some 6 years ago. The team of Chuck Murry at the University of Washington (UW) had spent almost two decades refining methods to grow large number of cardiomyocytes differentiated from human pluripotent stem cells (hPSC-CMs), and honing in the best way to deliver these to the injured myocardium so that they would survive, grow in size, and mature to become functional portions of a remuscolarized cardiac wall. The goal of moving to clinical trials had felt exceedingly close, until in the early '2010s studies in large animal models came as a wet blanket by revealing the high prevalence of engraftment-associated arrhythmias (EAs). This was a major roadblock that required an "all hands on deck" approach. The predominant hypothesis was that the immaturity of hPSC-CMs was to blame, leading them to behave as an ectopic "pacemaker". Many approaches had been tested to improve the maturity of hPSC-CMs, such as mechanical, hormonal, or metabolic stimulation. All had provided some important advances, but none had yet fully succeeded.
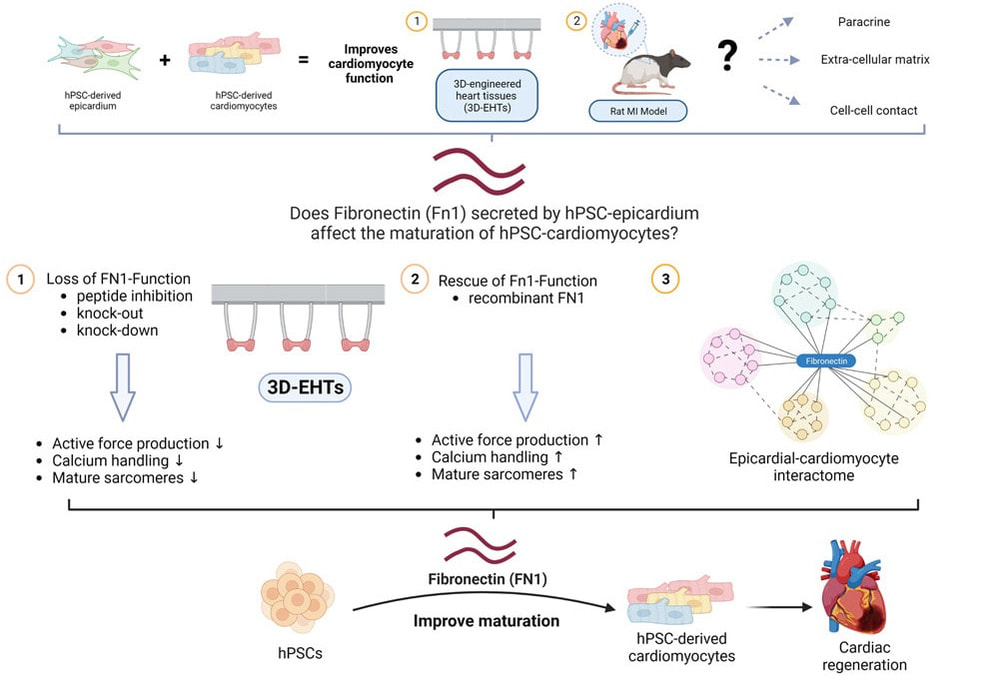
Desperate times call for desperate measures, and so Chuck decided it was time to try a drastic approach to eliminate the problem at its root: modify the genome to "educate" adolescent hPSC-CMs to behave like adults. He established a "Strikeforce" whose goal was to identify the right targets, edit them in or out of the genome of hPSCs, and test the resulting hPSC-CMs for their ability to spontaneously beat in vitro, a possible surrogate of their arrhythmogenic properties in vivo. When I had joined Chuck's lab as a postdoc a couple of years prior it was for the 4D Nucleome project, a basic science effort to discover the functional changes in 3D chromatin structure in cardiac development and disease. Since then, I had grown increasingly interested in the efforts of the Heart Regeneration Project (HRP), but I could not see how to contribute given my background in gene regulation and genome editing. Until one day, during a long walk along Lake Union, Chuck asked me if I wanted to lead the Strikeforce. It was a fork in the road I took eagerly: the rest, as they say, is history. The MEDUSA project was the most ambitious I have ever been part of; it thought me patience, resilience, leadership, and the true meaning of teamwork. Besides the Strikeforce's members, first and foremost lead author Silvia Marchianò, some 25 scientists across academia and industry were all pivotal to produce cells in large scale, transplant them, and study the resulting animals and tissues. What was supposed to be a quick "in and out" operation turned into a gruelling war of attrition with an enemy that proved crafty and full of resources. After generating some 12 gene edited lines, we finally identified a combination of 4 genome edits able to *finally* prevent spontaneous beating in hPSC-CMs in vitro, yet maintain their ability to respond to an electrical signal. In agreement with our initial hypothesis, their transplantation did not result in EA. Following the footsteps of Perseus, we had finally slayed the MEDUSA. I left the UW to start my lab in Italy two years ago almost exactly, and since then the study continued, marching through a though but fair peer review, among other things confirming our initial safety findings also when a larger dose of hPSC-CMs were transplanted. I cannot thank Chuck enough for his steadfast support of the study even after the many failures at the beginning. The Strikeforce and HRP as a whole are the inspiration for the program I am working hard to establish, with different goals, also on our side of the pond. The mythos does not end here: there are still several monsters to be slain on the path towards remuscolarizing the heart of patients in a safe, efficient, and scalable manner. Still, today we stop and celebrate that we are, at last, a big step closer. Alessandro Bertero Transatlantic collaboration identifies key molecule for cardiac bioengineering (and regeneration?)3/30/2023 Our friends and collaborators in the Sinha lab in Cambridge published an exciting paper in Stem Cell Reports that reports the key role of an extracellular matrix component, fibronectin, in the maturation of cardiomyocytes seeded in 3D scaffolds together with epicardial cells (forming so-called 3D Engineered Heart Tissues, 3D-EHTs). The study involved a transatlantic collaboration with the Murry lab in Seattle, WA. We were glad to contribute to the genome editing experiments that led to the formal demonstration that fibronectin is required for the pro-maturation activity of epicardial dells.
A previous study from the same collaborative group had identified they key ability of epicardial cells to promote cardiomyocyte maturation both in vitro and in vivo, but the mechanism remained unclear. This study identifies fibronecting as being both sufficient and necessary for this process in vitro, which opens the door to simplified and more reproducible bioengineering applications compared to adding epicardial cells.
Fibronectin may prove key to also boost cardiac remuscolarization in vivo. Indeed, the immaturity of human pluripotent-derived cardiomyocytes (hPSC-CMs) is thought to be a key driver of arrhythmic complications following their transplantation: what if a pinch of fibronectin in transplanted hPSC-CMs improved their safe engraftment and function?There promises to be a third episode of this saga.. Paper sheds lights on chromatin topology dynamics during endothelial vs. cardiac differentiation12/12/2022 A novel study we contributed to provides the first detailed analysis of 3D chromatin organisation changes during the development of endothelium, and how they correlate with gene expression regulations. The study also compares these dynamics with those we previously reported for cardiomyogenesis; here are the highlights:
This work was led by two talented postdoc in the Murry and Noble laboratories at the University of Washington, Katie Mitzelfelt and Kris Alavattam, and was published on the ISSCR journal Stem Cell Report: https://www.cell.com/stem-cell-reports/fulltext/S2213-6711(22)00536-7 We are honoured to be among the FEBS Excellence Awardees class of '22!
FEBS launched a prestigious new programme in 2021 aimed at supporting highly competitive research in molecular life sciences: the FEBS Excellence Awards. The FEBS Excellence Awards provide €100,000 funding over three years to early-career group leaders working in a FEBS country to purchase laboratory equipment and consumables. We are extremely grateful to FEBS for the support and we are eager to put these funds to good use to support our project entitled: "Chromatin Topology Dysregulation in Cardiac Cohesinopathies" We are particularly proud to be the first group receiving this award in Italy, and we would be more than happy to help any perspective applicant in preparing their application. The call for applications for FEBS Excellence Awards in 2023 is expected to open in April 2023. The closing date will be 1 July 2023. More info and full list of awardees on the FEBS website: https://www.febs.org/news/febs-excellence-awardees-2022/ |